Click to show/hide the code
pacman::p_load(tidyverse, ggstatsplot)Lim Li Ying
In this exercise, other than tidyverse, ggstatsplot will also be used. ggstatsplot is an extension of the ggplot2 package and allows details from statistical tests to be added to the information-rich plots.
Rows: 322 Columns: 7
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (4): ID, CLASS, GENDER, RACE
dbl (3): ENGLISH, MATHS, SCIENCE
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.# A tibble: 322 × 7
ID CLASS GENDER RACE ENGLISH MATHS SCIENCE
<chr> <chr> <chr> <chr> <dbl> <dbl> <dbl>
1 Student321 3I Male Malay 21 9 15
2 Student305 3I Female Malay 24 22 16
3 Student289 3H Male Chinese 26 16 16
4 Student227 3F Male Chinese 27 77 31
5 Student318 3I Male Malay 27 11 25
6 Student306 3I Female Malay 31 16 16
7 Student313 3I Male Chinese 31 21 25
8 Student316 3I Male Malay 31 18 27
9 Student312 3I Male Malay 33 19 15
10 Student297 3H Male Indian 34 49 37
# ℹ 312 more rowsgghistostats()The code chunk below uses gghistostats() to create a visual of a one-sample test on English scores.
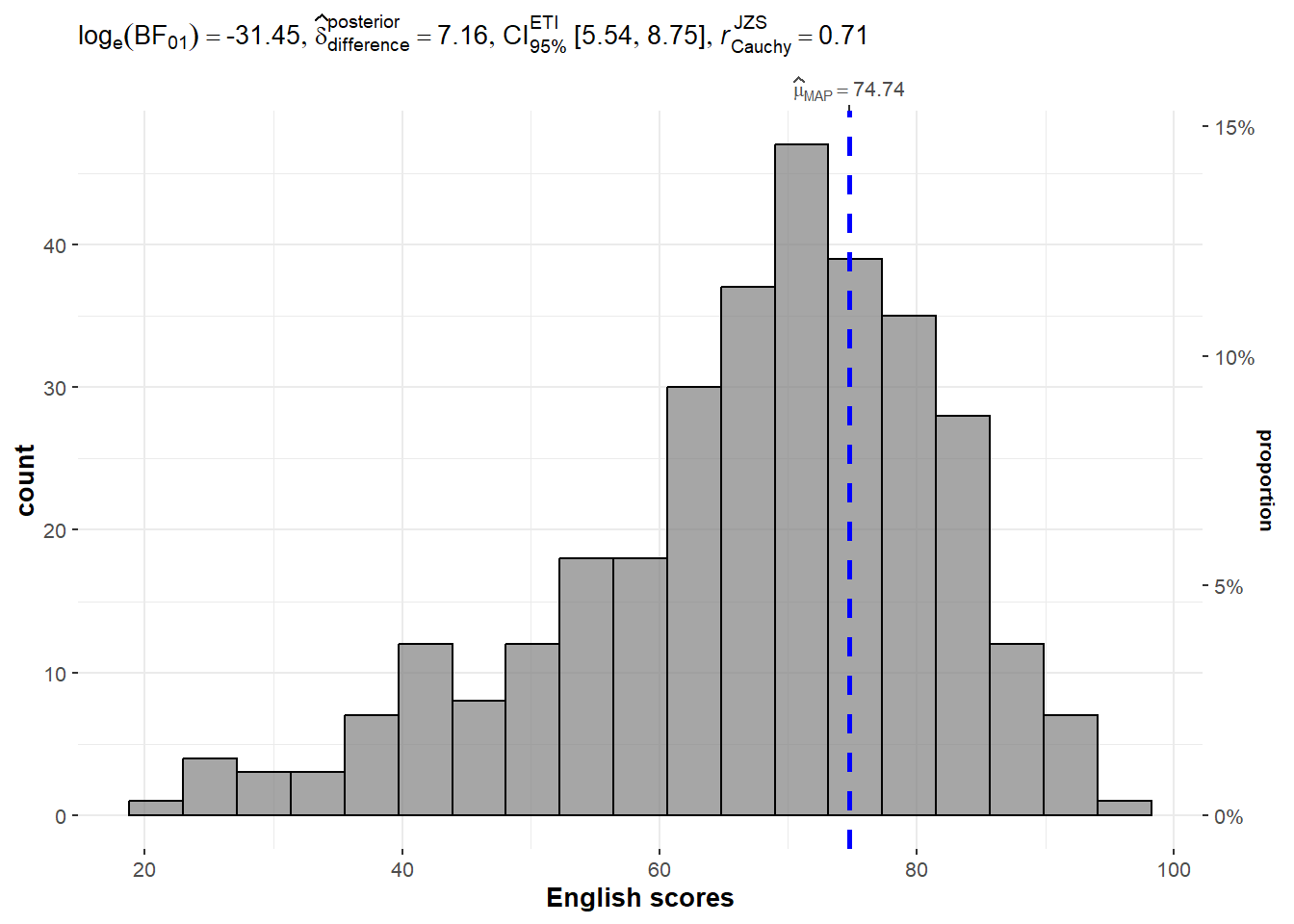
Set seed is important especially when doing variance statistics in order to ensure reproducibility.
ggbetweenstats()The code chunk below uses ggbetweenstats() to create a visual of a two-sample mean test of Maths scores by gender.
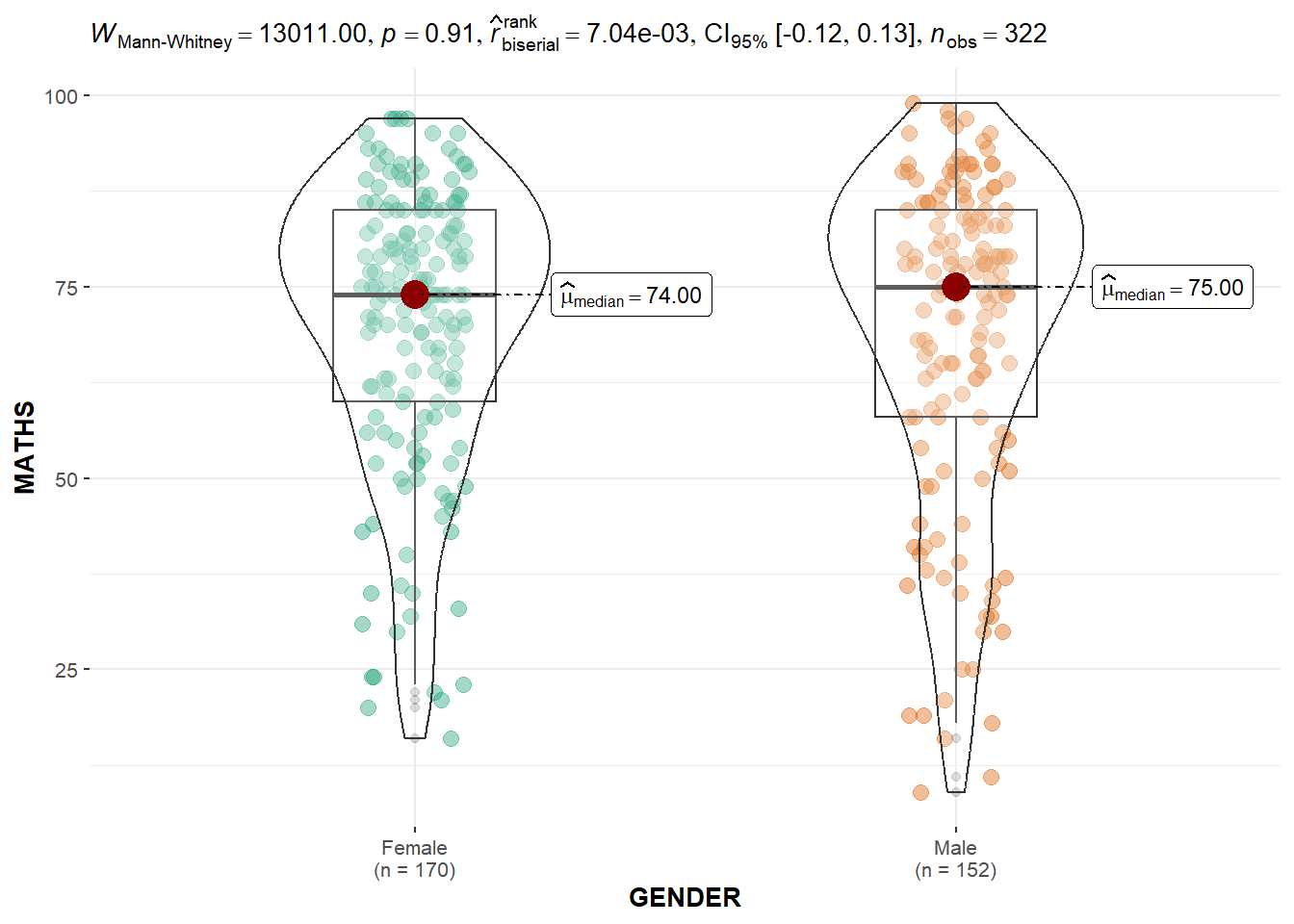
There are 4 types of statistical approach:
p <- parametric
np <- nonparametric
r <- robust
b <- bayes
ggbetweenstats()The code chunk below uses ggbetweenstats() to create a visual of a oneway ANOVA test of English scores by race.
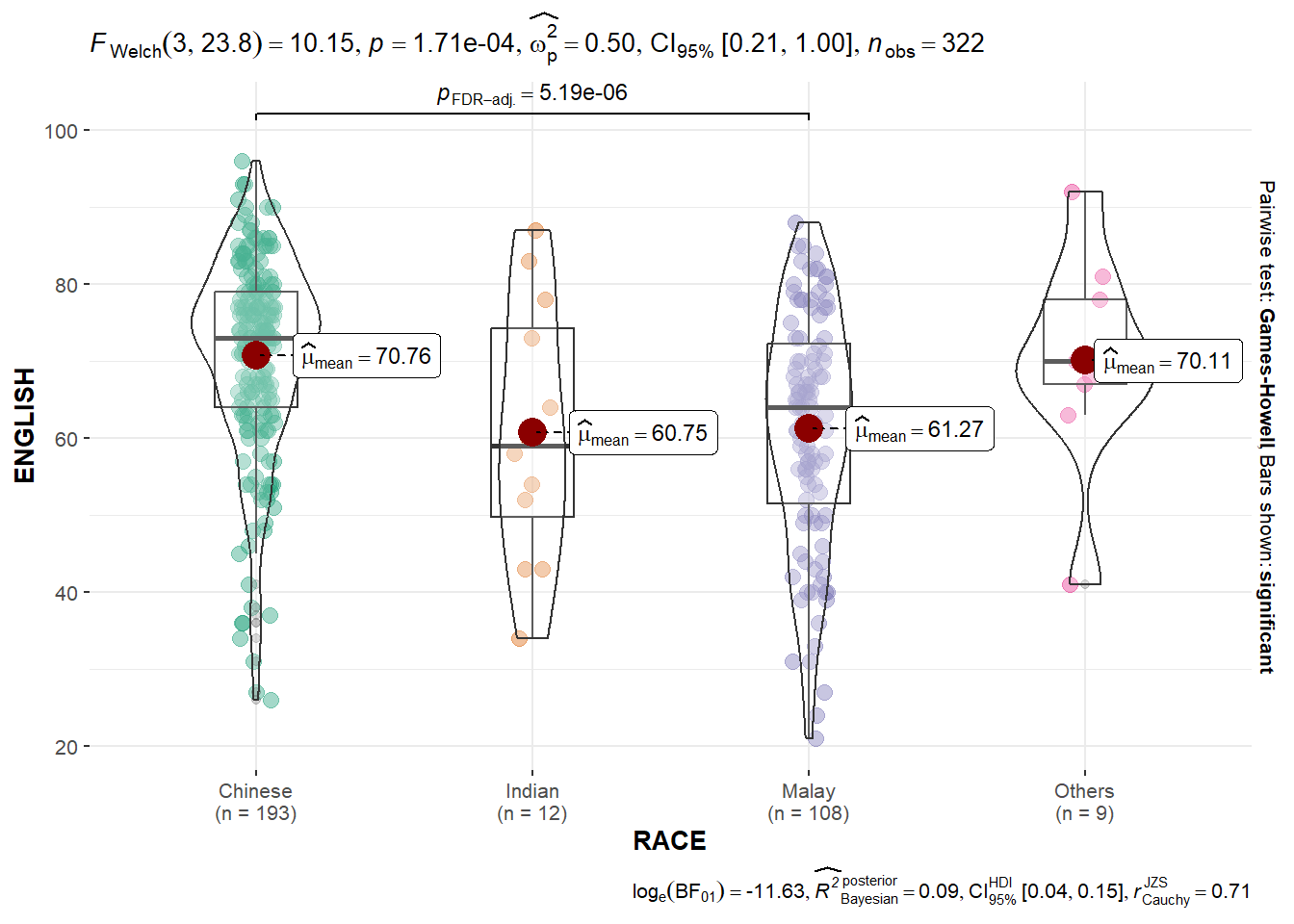
For pairwise display,
s <- displays only significant pairwise comparisons
ns <- displays only non-significant pairwise comparisons
all <- displays all pairwise comparisons
ggscatterstats()The code chunk below uses ggscatterstats() to create a visual of a Significant Test of Correlation between Maths and English scores.
ggbarstats()Firstly, the Maths scores are binned into 4 groups using cut().
The code chunk below uses ggbarstats() to create a visual of a Significant Test of Association between Maths scores and gender.
The code chunk below shows the R packages to be installed and loaded into the R environment for this exercise.
To import the excel worksheet ToyotaCorolla.xls, read_xls() of the readxl package is used, as seen in the code chunk below.
# A tibble: 1,436 × 38
Id Model Price Age_08_04 Mfg_Month Mfg_Year KM Quarterly_Tax Weight
<dbl> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 81 TOYOTA … 18950 25 8 2002 20019 100 1180
2 1 TOYOTA … 13500 23 10 2002 46986 210 1165
3 2 TOYOTA … 13750 23 10 2002 72937 210 1165
4 3 TOYOTA… 13950 24 9 2002 41711 210 1165
5 4 TOYOTA … 14950 26 7 2002 48000 210 1165
6 5 TOYOTA … 13750 30 3 2002 38500 210 1170
7 6 TOYOTA … 12950 32 1 2002 61000 210 1170
8 7 TOYOTA… 16900 27 6 2002 94612 210 1245
9 8 TOYOTA … 18600 30 3 2002 75889 210 1245
10 44 TOYOTA … 16950 27 6 2002 110404 234 1255
# ℹ 1,426 more rows
# ℹ 29 more variables: Guarantee_Period <dbl>, HP_Bin <chr>, CC_bin <chr>,
# Doors <dbl>, Gears <dbl>, Cylinders <dbl>, Fuel_Type <chr>, Color <chr>,
# Met_Color <dbl>, Automatic <dbl>, Mfr_Guarantee <dbl>,
# BOVAG_Guarantee <dbl>, ABS <dbl>, Airbag_1 <dbl>, Airbag_2 <dbl>,
# Airco <dbl>, Automatic_airco <dbl>, Boardcomputer <dbl>, CD_Player <dbl>,
# Central_Lock <dbl>, Powered_Windows <dbl>, Power_Steering <dbl>, …lm()The following code chunk is uses lm() to build a multiple linear regression model.
Call:
lm(formula = Price ~ Age_08_04 + Mfg_Year + KM + Weight + Guarantee_Period,
data = car_resale)
Coefficients:
(Intercept) Age_08_04 Mfg_Year KM
-2.637e+06 -1.409e+01 1.315e+03 -2.323e-02
Weight Guarantee_Period
1.903e+01 2.770e+01 Multicollinearity can be checked using check_collinearity() of the performance package, as seen in the code chunk below.
# Check for Multicollinearity
Low Correlation
Term VIF VIF 95% CI Increased SE Tolerance Tolerance 95% CI
KM 1.46 [ 1.37, 1.57] 1.21 0.68 [0.64, 0.73]
Weight 1.41 [ 1.32, 1.51] 1.19 0.71 [0.66, 0.76]
Guarantee_Period 1.04 [ 1.01, 1.17] 1.02 0.97 [0.86, 0.99]
High Correlation
Term VIF VIF 95% CI Increased SE Tolerance Tolerance 95% CI
Age_08_04 31.07 [28.08, 34.38] 5.57 0.03 [0.03, 0.04]
Mfg_Year 31.16 [28.16, 34.48] 5.58 0.03 [0.03, 0.04]Variables with a Variance Inflation Factor (VIF) score of >10 are considered to be highly correlated. In this case, age and manufacturing year have high correlation. Therefore one of the variables will need to be removed from subsequent models.
The normality assumption can be checked using check_normality() of the performance package, as seen in the code chunk below.
Homogeneity of variances can be checked using check_heteroscedasticity() of the performance package, as seen in the code chunk below.
We can also perform a complete check using check_model().
ggcoefstats()---
title: "Hands-on Exercise 4"
author: "Lim Li Ying"
---
# *Visual Statistical Analysis with ggstatsplot*
# Getting Started
## Installing and loading R packages
In this exercise, other than *tidyverse*, *ggstatsplot* will also be used. *ggstatsplot* is an extension of the *ggplot2* package and allows details from statistical tests to be added to the information-rich plots.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
pacman::p_load(tidyverse, ggstatsplot)
```
## Loading the data
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
exam_data <- read_csv("data/Exam_data.csv")
exam_data
```
# One-sample test using `gghistostats()`
The code chunk below uses `gghistostats()` to create a visual of a one-sample test on English scores.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
set.seed(1234)
gghistostats(
data = exam_data,
x = ENGLISH,
type = "bayes",
test.value = 60,
xlab = "English scores"
)
```
::: callout-note
Set seed is important especially when doing variance statistics in order to ensure reproducibility.
:::
# Two-sample mean test using `ggbetweenstats()`
The code chunk below uses `ggbetweenstats()` to create a visual of a two-sample mean test of Maths scores by gender.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
ggbetweenstats(
data = exam_data,
x = GENDER,
y = MATHS,
type = "np",
messages = FALSE
)
```
::: callout-note
There are 4 types of statistical approach:
1. `p` \<- parametric
2. `np` \<- nonparametric
3. `r` \<- robust
4. `b` \<- bayes
:::
# Oneway ANOVA test using `ggbetweenstats()`
The code chunk below uses `ggbetweenstats()` to create a visual of a oneway ANOVA test of English scores by race.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
ggbetweenstats(
data = exam_data,
x = RACE,
y = ENGLISH,
type = "p",
mean.ci = TRUE,
pairwise.comparisons = TRUE,
pairwise.display = "s",
p.adjust.method = "fdr",
messages = FALSE
)
```
::: callout-note
For pairwise display,
- `s` \<- displays only significant pairwise comparisons
- `ns` \<- displays only non-significant pairwise comparisons
- `all` \<- displays all pairwise comparisons
:::
# Significant test of correlation using `ggscatterstats()`
The code chunk below uses `ggscatterstats()` to create a visual of a Significant Test of Correlation between Maths and English scores.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
ggscatterstats(
data = exam_data,
x = MATHS,
y = ENGLISH,
marginal = FALSE
)
```
# Significant test of association (dependence) using `ggbarstats()`
Firstly, the Maths scores are binned into 4 groups using `cut()`.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
exam_data1 <- exam_data %>%
mutate(MATHS_bins =
cut(MATHS, breaks = c(0,60,75,85,100))
)
```
The code chunk below uses `ggbarstats()` to create a visual of a Significant Test of Association between Maths scores and gender.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
ggbarstats(
data = exam_data1,
x = MATHS_bins,
y = GENDER
)
```
# *Visualizing Models*
# Getting Started
## Installing and loading R packages
The code chunk below shows the R packages to be installed and loaded into the R environment for this exercise.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
pacman::p_load(readxl, performance, parameters, see)
```
## Loading the data
To import the excel worksheet *ToyotaCorolla.xls*, `read_xls()` of the *readxl* package is used, as seen in the code chunk below.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
car_resale <- read_xls("data/ToyotaCorolla.xls")
car_resale
```
# Multiple linear regression modelling using `lm()`
The following code chunk is uses `lm()` to build a multiple linear regression model.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
model <- lm(Price ~ Age_08_04 + Mfg_Year + KM +
Weight + Guarantee_Period, data = car_resale)
model
```
# Model diagnostics: checking for multicollinearity
Multicollinearity can be checked using `check_collinearity()` of the *performance* package, as seen in the code chunk below.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
check_collinearity(model)
```
::: callout-note
Variables with a Variance Inflation Factor (VIF) score of \>10 are considered to be highly correlated. In this case, age and manufacturing year have high correlation. Therefore one of the variables will need to be removed from subsequent models.
:::
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
check_c <- check_collinearity(model)
plot(check_c)
```
# Model diagnostic: checking normality assumption
The normality assumption can be checked using `check_normality()` of the *performance* package, as seen in the code chunk below.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
# build another model after removing "Mfg_Year"
model1 <- lm(Price ~ Age_08_04 + KM +
Weight + Guarantee_Period, data = car_resale)
check_n <- check_normality(model1)
plot(check_n)
```
# Model diagnostic: checking for homogeneity of variances
Homogeneity of variances can be checked using `check_heteroscedasticity()` of the *performance* package, as seen in the code chunk below.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
check_h <- check_heteroscedasticity(model1)
plot(check_h)
```
# Model diagnostic: complete check
We can also perform a complete check using `check_model()`.
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
check_model(model1)
```
# Visualizing regression parameters using *see*
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
plot(parameters(model1))
```
# Visualizing regression parameters using `ggcoefstats()`
```{r}
#| code-fold: show
#| code-summary: "Click to show/hide the code"
ggcoefstats(model1,
output = "plot")
```